- java.lang.Object
-
- io.jenetics.ProbabilitySelector<G,C>
-
- io.jenetics.LinearRankSelector<G,C>
-
- All Implemented Interfaces:
- Selector<G,C>
public final class LinearRankSelector<G extends Gene<?,G>,C extends Comparable<? super C>> extends ProbabilitySelector<G,C>
In linear-ranking selection the individuals are sorted according to their fitness values. The rank N is assignee to the best individual and the rank 1 to the worst individual. The selection probability P(i) of individual i is linearly assigned to the individuals according to their rank.
Here n-/N is the probability of the worst individual to be selected and n+/N the probability of the best individual to be selected. As the population size is held constant, the conditions n+ = 2 - n- and n- >= 0 must be fulfilled. Note that all individuals get a different rank, i.e., a different selection probability, even if the have the same fitness value.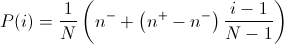
T. Blickle, L. Thiele, A comparison of selection schemes used in evolutionary algorithms, Technical Report, ETH Zurich, 1997, page 37. http://citeseer.ist.psu.edu/blickle97comparison.html
- Since:
- 1.0
- Version:
- 5.0
- Author:
- Franz Wilhelmstötter
-
-
Field Summary
-
Fields inherited from class io.jenetics.ProbabilitySelector
_reverter, _sorted, POPULATION_COMPARATOR
-
-
Constructor Summary
Constructors Constructor and Description LinearRankSelector()Create a new LinearRankSelector withnminus := 0.5.LinearRankSelector(double nminus)Create a new LinearRankSelector with the given values fornminus.
-
Method Summary
All Methods Instance Methods Concrete Methods Modifier and Type Method and Description protected double[]probabilities(Seq<Phenotype<G,C>> population, int count)This method sorts the population in descending order while calculating the selection probabilities.StringtoString()-
Methods inherited from class io.jenetics.ProbabilitySelector
probabilities, select
-
-
-
-
Constructor Detail
-
LinearRankSelector
public LinearRankSelector(double nminus)
Create a new LinearRankSelector with the given values fornminus.- Parameters:
nminus-nminus/Nis the probability of the worst phenotype to be selected.- Throws:
IllegalArgumentException- ifnminus < 0.
-
LinearRankSelector
public LinearRankSelector()
Create a new LinearRankSelector withnminus := 0.5.
-
-
Method Detail
-
probabilities
protected double[] probabilities(Seq<Phenotype<G,C>> population, int count)
This method sorts the population in descending order while calculating the selection probabilities.- Specified by:
probabilitiesin classProbabilitySelector<G extends Gene<?,G>,C extends Comparable<? super C>>- Parameters:
population- The unsorted population.count- The number of phenotypes to select. This parameter is not needed for most implementations.- Returns:
- Probability array. The returned probability array must have the
length
population.size()and must sum to one. The returned value is checked withassert(Math.abs(math.sum(probabilities) - 1.0) < 0.0001)in the base class.
-
-